PalmLab Help Center
Search Function
Search for specific proteins and palmitoylation sites across the entire PalmLab database.
1 Search Interface
The search page provides multiple ways to find proteins:
Search Options:
- Search Box: Enter any search term
- Search Type:
- All Fields (default)
- UniProt ID
- Gene Symbol
- Protein Name
- Organism Filter:
- All Organisms
- Human only
- Mouse only
Input Examples:
Gene Symbols: KRAS, TP53, BRAF
Protein Names: "GTPase KRas", "Cellular tumor antigen p53"
Mixed Input: "P01116 KRAS TP53"
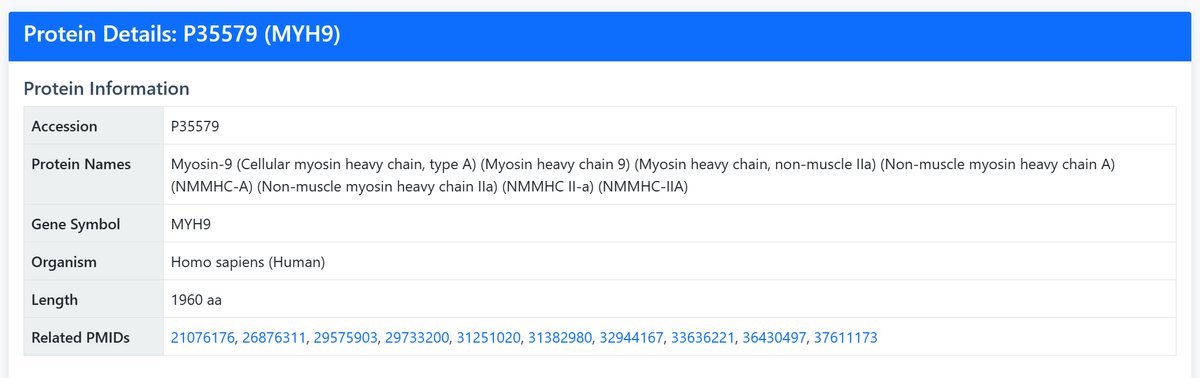
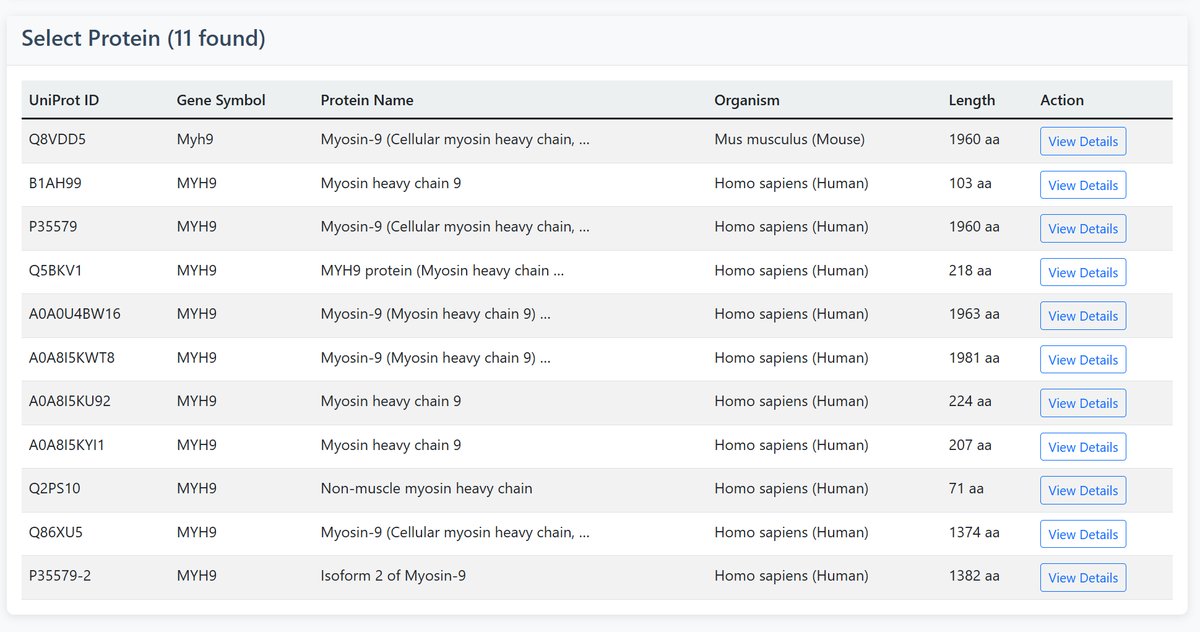
If the search yields multiple results, you can click “View Details” for the target protein in the table to examine it.
2 Search Results
After searching, you'll see a results table with the following information:
| Column | Description | Example |
|---|---|---|
| UniProt ID | Unique protein identifier | P35579 |
| Gene Symbol | Standard gene name | MYH9 |
| Protein Name | Full protein description | Myosin-9 |
| Organism | Species information | Homo sapiens |
| Length | Protein length in amino acids | 1960 aa |
| Action | Link to detailed view | "View Details" button |
3 Protein Details Page
Click "View Details" to access comprehensive protein information:
Protein Information Section:
- Accession: UniProt identifier
- Protein Names: Full descriptive name
- Gene Symbol: Standard gene name
- Organism: Species information
- Length: Protein size in amino acids
- Related PMIDs: Links to relevant publications

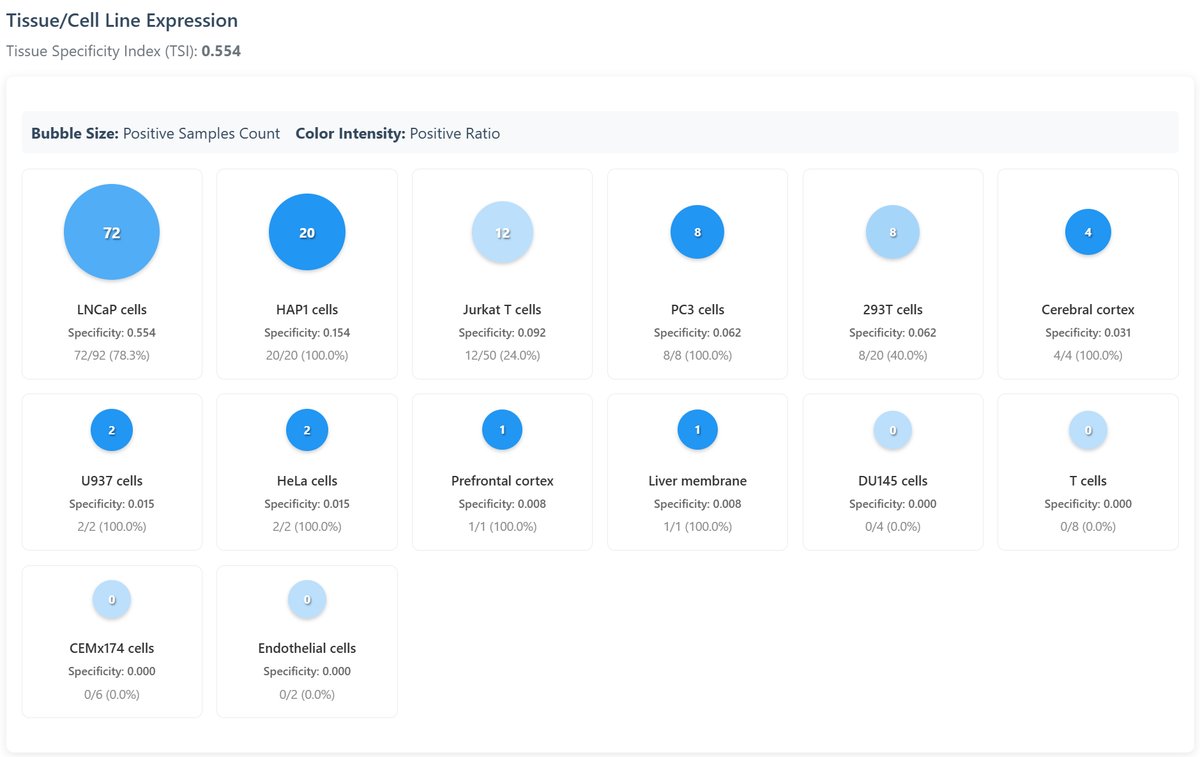
Tissue/Cell Line Expression:
Visual representation of palmitoylation across different tissues:
- Bubble Heatmap: Show the quantity and proportion of palmitoylated proteins found in all the tissues and cell lines in the database.
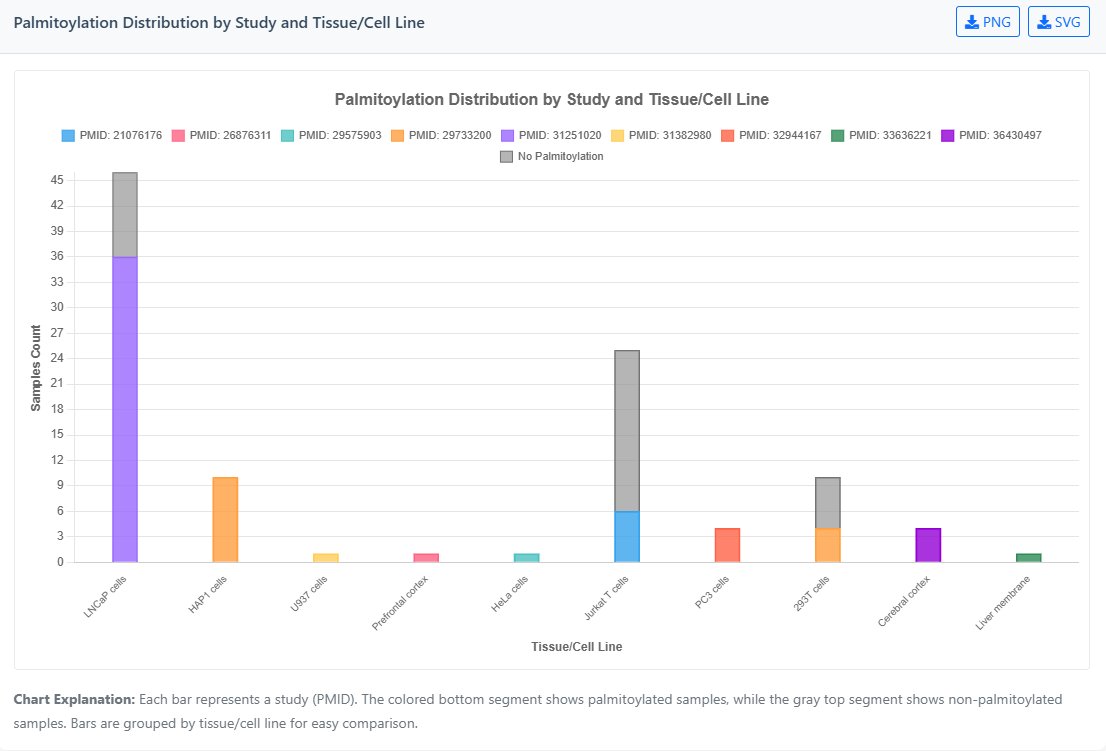
- Bar Chart: This stacked bar chart displays palmitoylation detection across different research studies (PMID). Coloured segments represent positive samples, while grey segments indicate negative samples from each publication.
- TSI Index: Tissue Specificity Index (higher = more specific) TSI = 0: Ubiquitous expression TSI = 1: Highly tissue-specific"
Protein Sequence with Site Annotation:
The sequence display uses color-coded highlighting:
Single Types:
Combined Types:
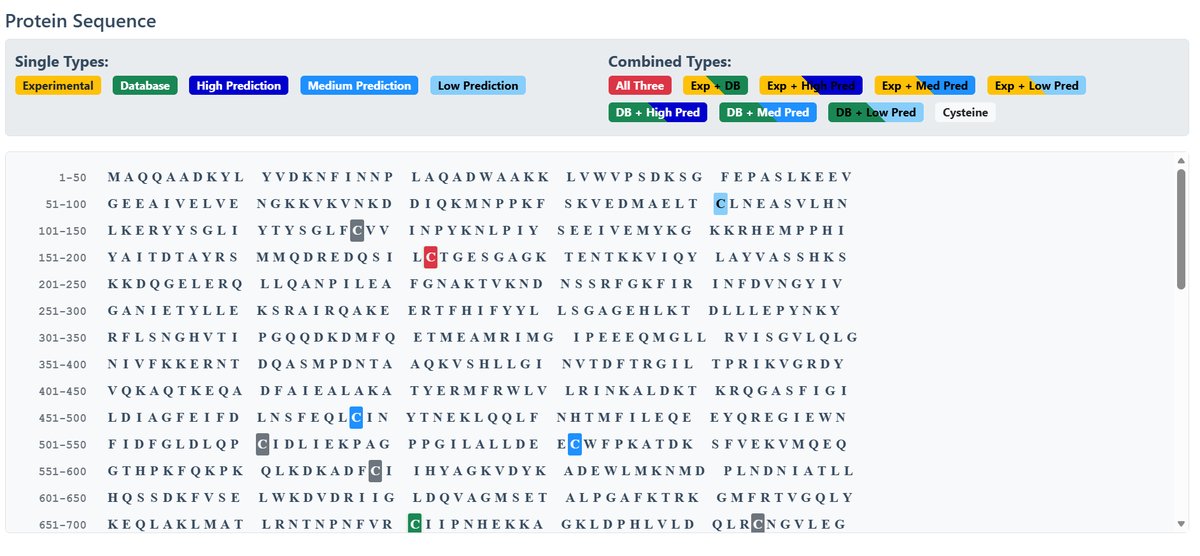
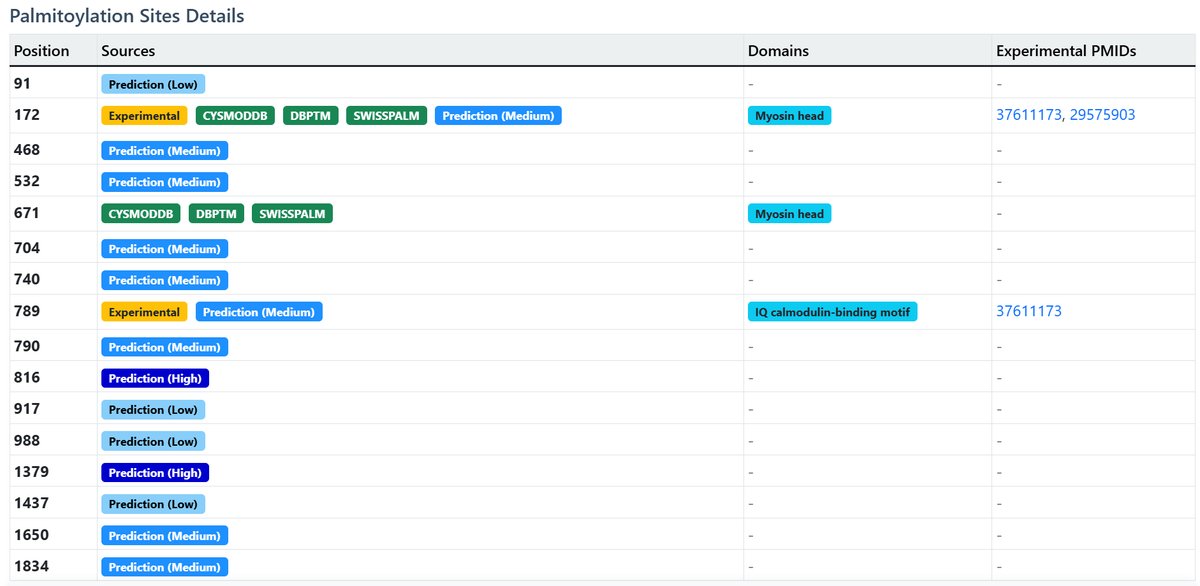
Palmitoylation Sites Details:
Comprehensive table of all known palmitoylation sites:
| Position | Sources | Domain | Experimental PMIDs |
|---|---|---|---|
| 91 | Prediction (Low) | ||
| 172 | Experimental CysModDB dbPTM SwissPalm Prediction (High) | Myosin head | 37611173 29575903 |
PhyloP Conservation Scores:
Measures evolutionary conservation at individual bases using phylogenetic p-values:
- Positive scores: Indicate conservation (higher = more conserved)
- Negative scores: Indicate accelerated evolution (lower = faster evolution)
- Near zero: Neutral evolution
- Color coding: Blue shades for conservation, pink shades for acceleration
PhastCons Conservation Scores:
Identifies conserved elements using hidden Markov models:
- Range: 0 to 1
- Close to 1: Highly conserved, likely functional
- Close to 0: Fast evolving, less constrained
- Color coding: Blue shades for high conservation, pink shades for low conservation
Visualization Features:
- Three-base grouping: Each codon position shows scores for 1st, 2nd, and 3rd bases
- Interactive tooltips: Hover to see detailed scores and position information
- Position labels: Colored badges indicate site types (experimental, predicted, etc.)
- Scrollable view: Horizontal scrolling for proteins with many modification sites
- Human (hg38/GRCh38) Dataset:The “100 Vertebrates Alignment and Conservation (100-way)” dataset from the UCSC Genome Browser. This file contains whole-genome sequences from 100 vertebrate species aligned against the human hg38 reference genome.
- Mouse (mm39/GRCm39) Dataset:The “35 Vertebrates Alignment and Conservation (35-way)” dataset from the UCSC Genome Browser. This file contains whole-genome sequences from 35 vertebrate species aligned against the mouse mm39 reference genome.
Example Conservation Charts:
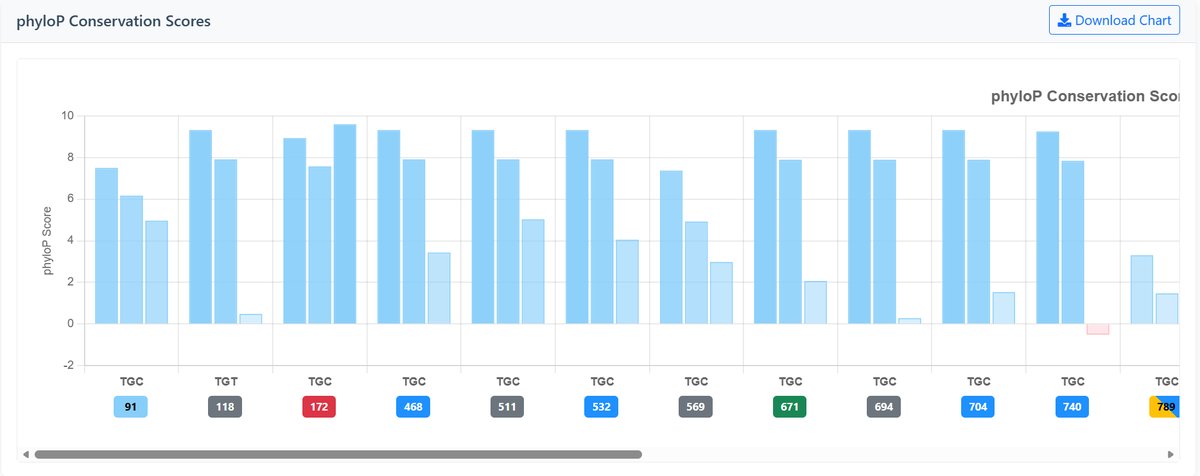
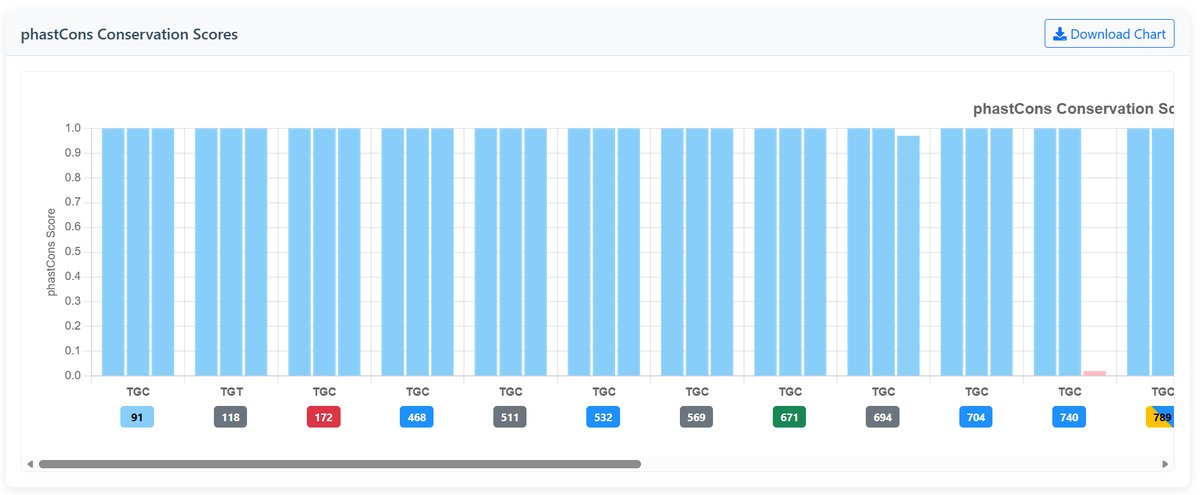
Interpretation Guide:
Highly conserved modification sites (high positive phyloP, phastCons close to 1) may indicate functionally important modifications.
Rapidly evolving sites (negative phyloP) may represent species-specific adaptations.
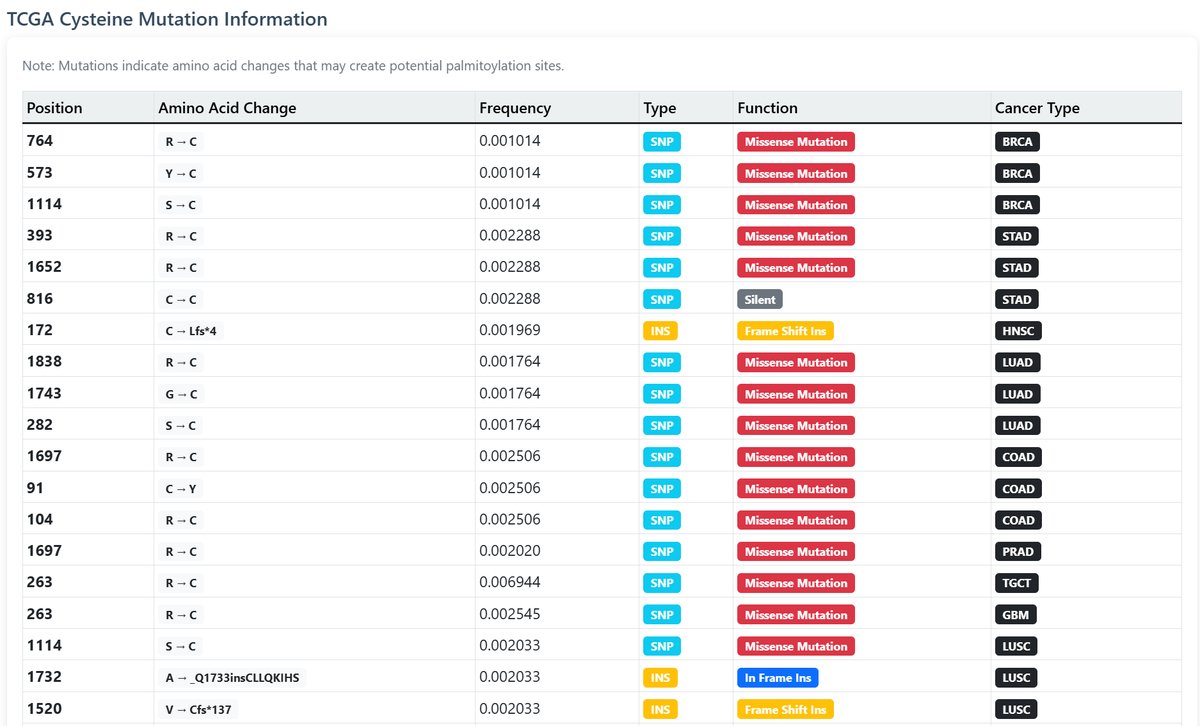
TCGA Mutation Information:
Cancer-related cysteine mutations that may create new palmitoylation sites:
- Mutation position and amino acid change
- Frequency in cancer samples
- Mutation type and functional impact
- Associated cancer types
Browse Database
Explore the PalmLab database through organized categories and filters.
1 By Database Source
Browse palmitoylated proteins by data source (top 100 from experimental studies displayed; all from database sources shown):
Available Sources:
- By Experiment: Proteins with experimental validation
- By SwissPalm: Curated data from SwissPalm database
- By CysModDB: Cysteine modification database entries
- By dbPTM: Database of Post-Translational Modifications
- By PTMD: PTM database entries
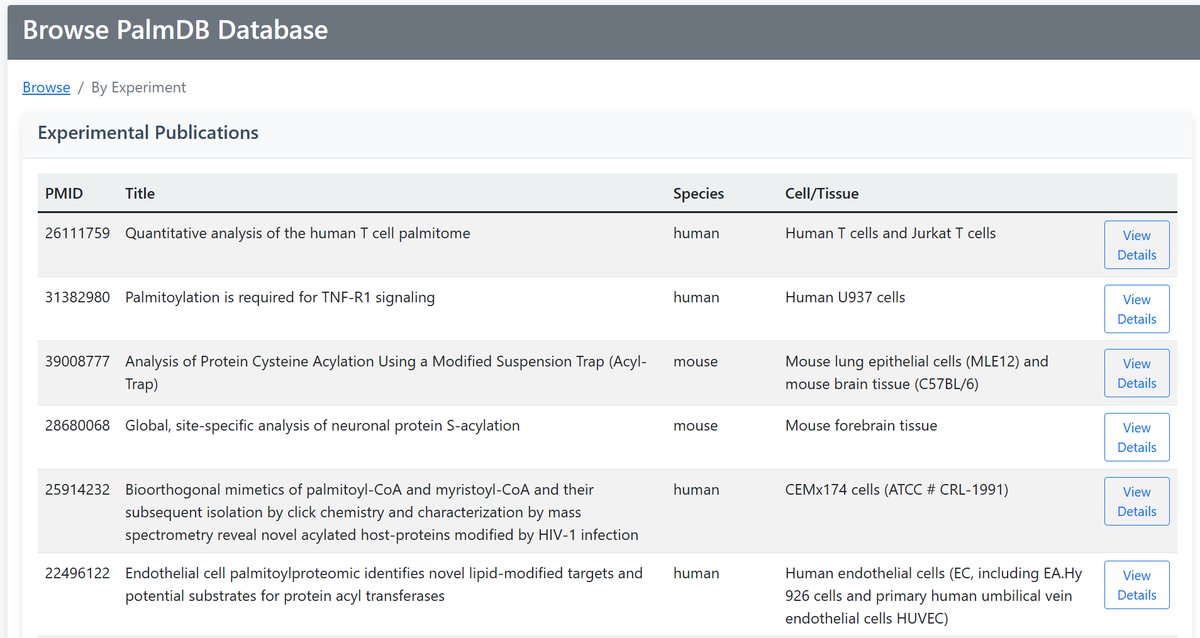
Example Output:
• PMID (Publication ID)
• Title
• Species
• Cell/Tissue
• Link to detailed view
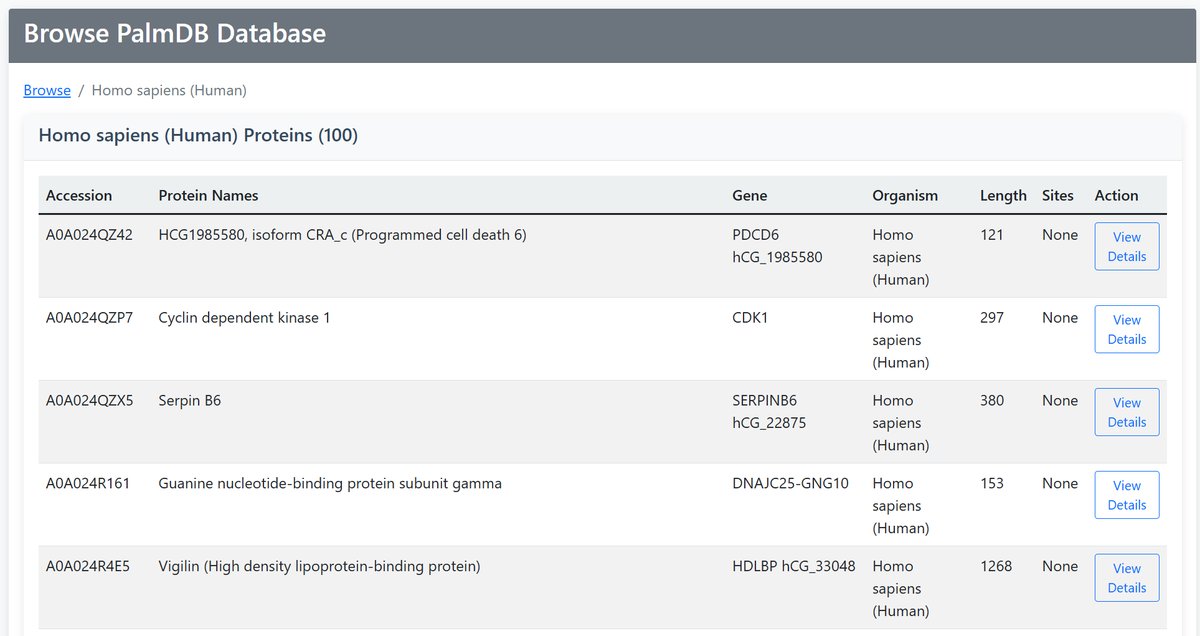
2 By Organism
Top 100 Proteins Browse by species:
Available Organisms:
- Homo sapiens (Human): Complete human proteome
- Mus musculus (Mouse): Mouse protein data
- All Organisms: View across all species
Organism Statistics:
Mouse: ~15,000 proteins
Total: ~35,000 proteins
Palmitoylation Sites: ~50,000 total
4 Browse Results
All browse views provide consistent protein information:
| Information | Description |
|---|---|
| Accession | UniProt identifier with link to details |
| Protein Names | Descriptive protein name |
| Gene | Gene symbol |
| Organism | Species information |
| Length | Protein size in amino acids |
| Sites | Number of palmitoylation sites |
| Action | "View Details" link to full protein page |
Tool 1: Differential Palmitoylation Analysis
Compare the palmitoylation status of query protein across different samples.
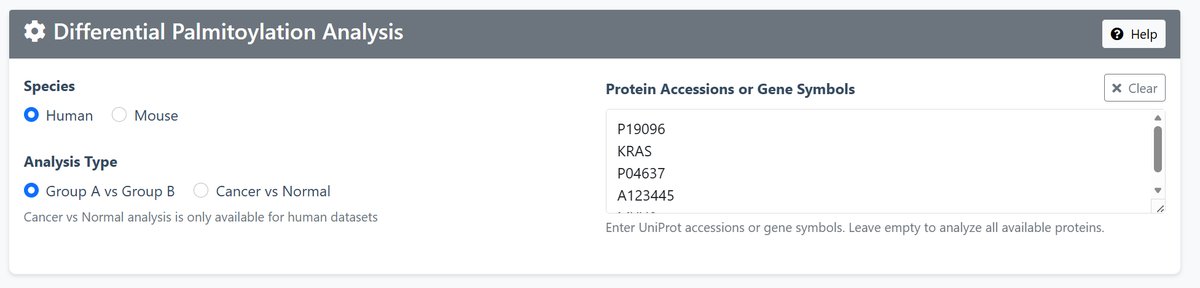
1 Configuration Setup
Species Selection: Choose between Human or Mouse data
Analysis Type:
- Group A vs Group B: Compare any two groups of datasets
- Cancer vs Normal: Specifically compare cancer vs normal tissues (Human only)
Protein Input: Enter UniProt accessions or gene symbols (e.g., P01116, KRAS, TP53)
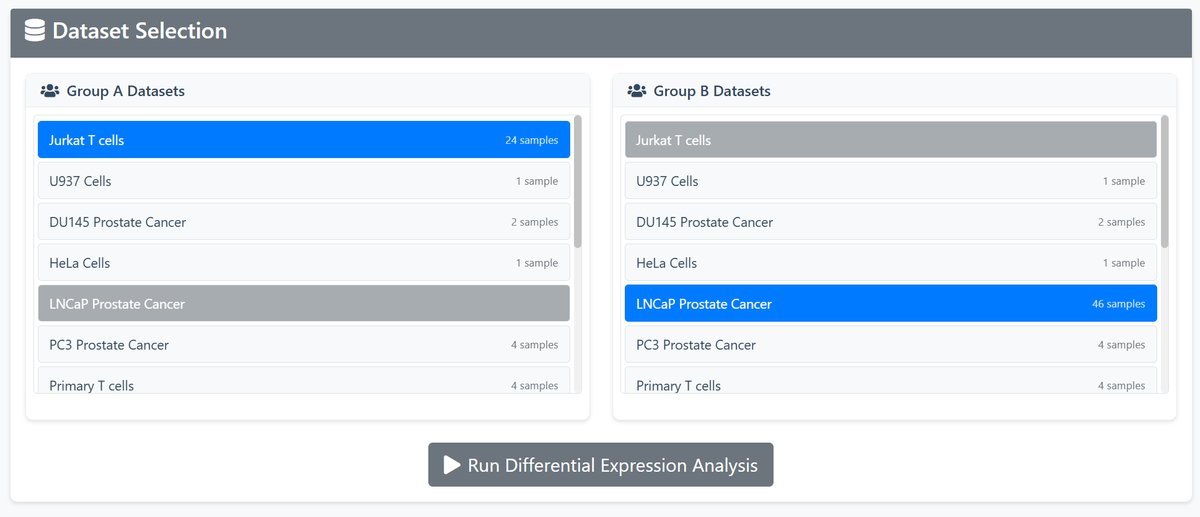
2 Dataset Selection
Available Datasets:
- Group A vs Group B: Compare any two groups of datasets
- Cancer vs Normal: Specifically compare cancer vs normal tissues (Human only)
3 Protein Validation
The system automatically validates input proteins and provides feedback:
- Not Found - Protein or gene not in database
- Species Mismatch - Protein or gene exists but in different species
- No Expression Data - Protein or gene is not expressed in either of the selected groups.
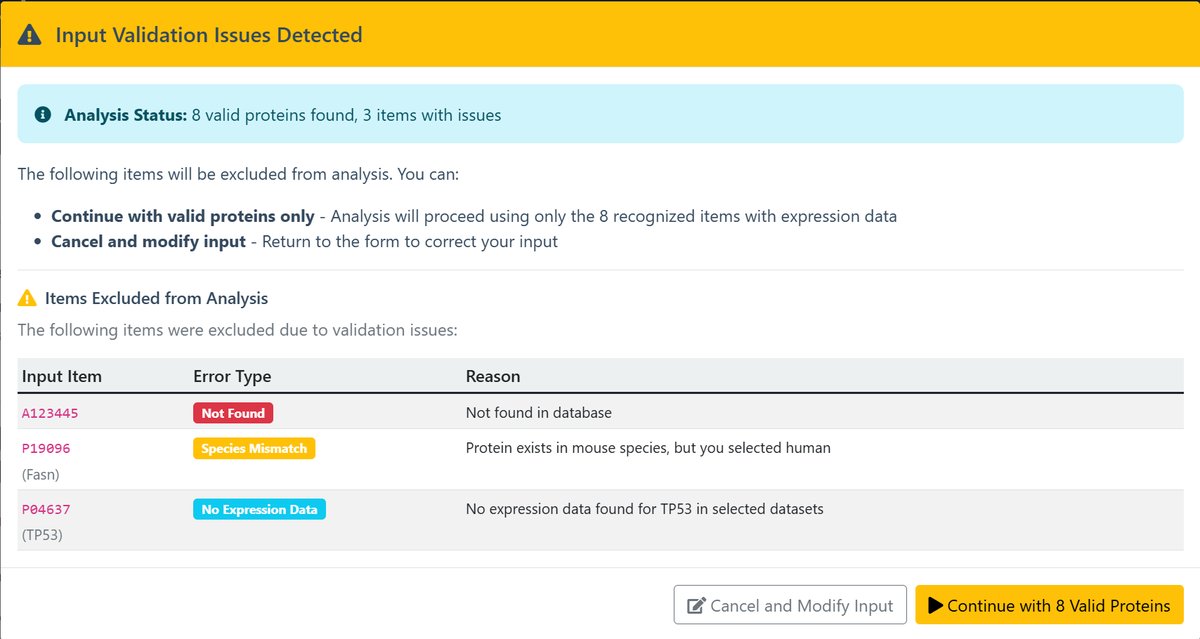
- Continue : Ignore these errors and proceed with analyzing the correct protein or gene.
- Cancel and Modify Input : Return to the previous step to make adjustments.
4 Analysis Execution
Click "Run Differential Expression Analysis" to start the analysis process.
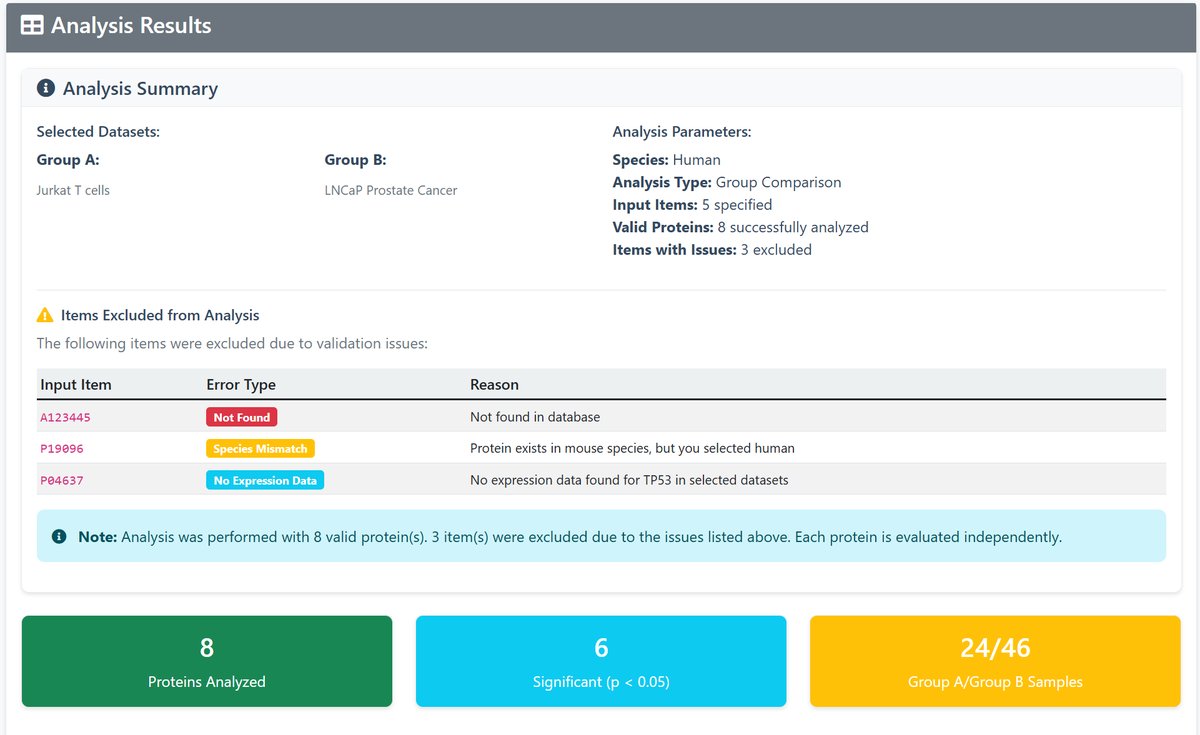
5 Results Interpretation
The results page provides:
- Summary Statistics: Overview of analyzed proteins and significant findings
- Results Table: Detailed comparison for each protein including:
- Protein accession and gene symbol
- Expression ratios for both groups
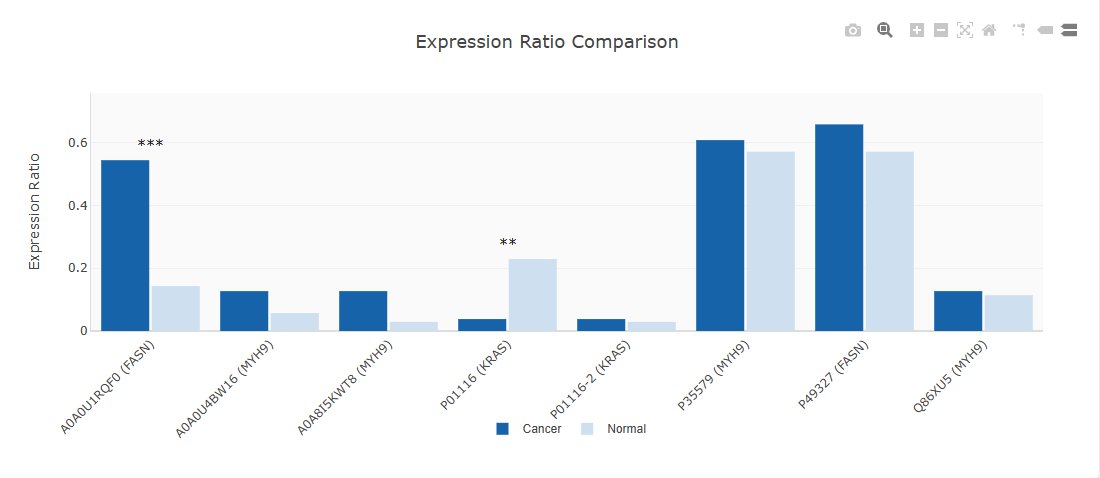
- Odds ratio and p-value
- Significance indicators (*p<0.05, **p<0.01, ***p<0.001)
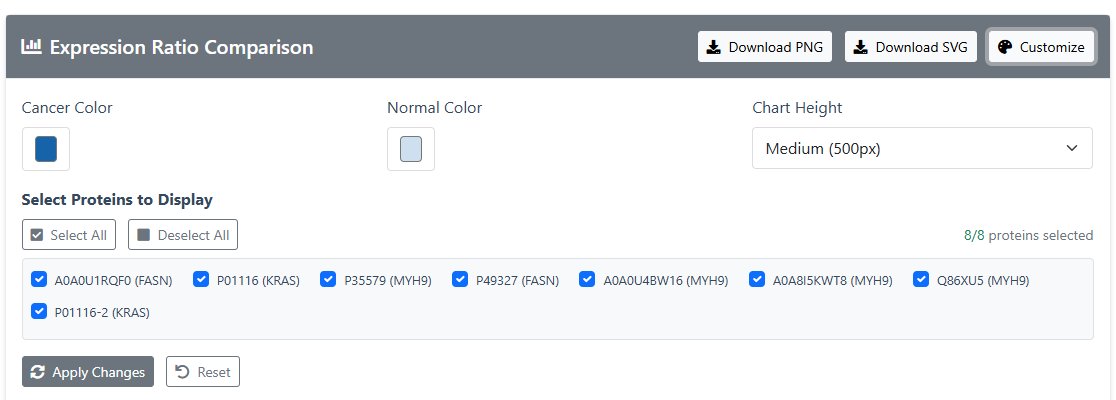
- Visualization: Interactive bar chart comparing expression ratios:
- Click "Customize"
- Customizable colors and selectable visual proteins
- Click “Apply Changes” to visualize.
- Visual images can be downloaded and saved by clicking “Download jpg” or “Download SVG”.
Tool 2: Palm-Protein Network Analysis
Found the exclusive or co-occurrent palmitoylation proteins of the query protein.
1 Input Configuration
Enter Query Protein: Input a Protein ID (e.g., P12345) or Gene Name (e.g., TP53)
Select Analysis Method:
- Fisher's Exact Test: Provides OR, P-value, FDR, and relationship type
- Jaccard Index: Provides Jaccard coefficient and relationship type
Choose Species: Human or Mouse
Select Tissue/Sample Type:
- Mouse: All, Liver, Brain
- Human: All, Tumor
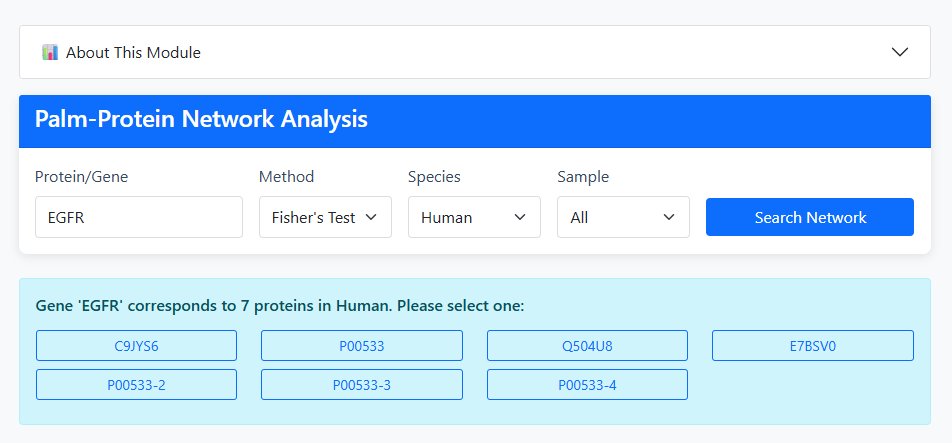
2 Analysis Execution
Click "Search Network" to execute the analysis and generate network visualization.
3 Results Interpretation
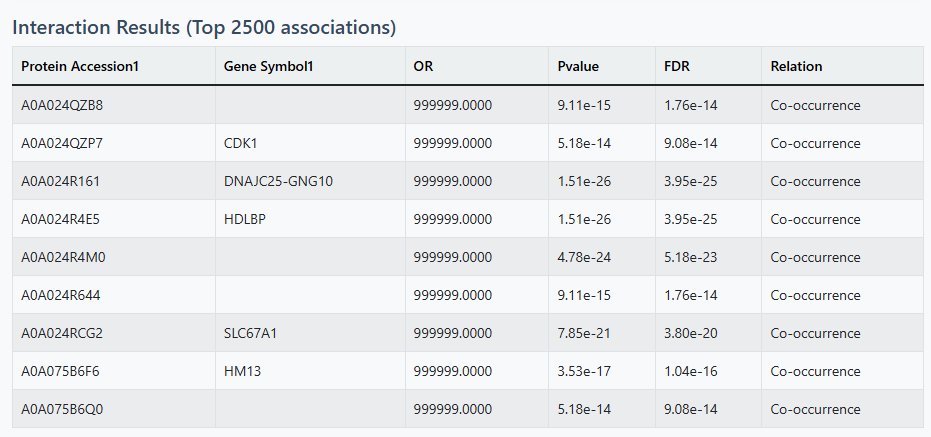
Table Results
- Top 2500 significant interactions displayed
- 25 results per page with pagination
- Columns include protein identifiers, association metrics, and relationship types
- Sortable by association strength (OR or Jaccard)
Network Visualization
- ● Core Protein (Query)
- ● First Level - Co-occurrence
- ● First Level - Mutual Exclusion
- ● Second Level - Co-occurrence
- ● Second Level - Mutual Exclusion
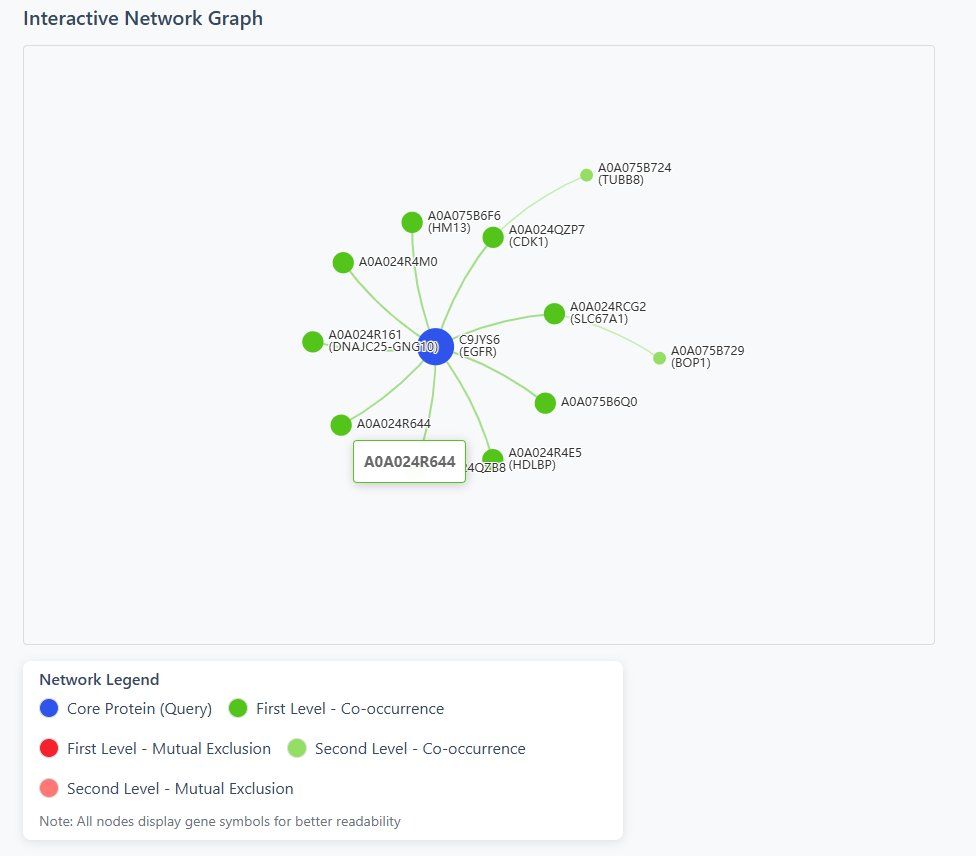
4 Interactive Features
- Drag nodes to rearrange network layout
- Hover over nodes for detailed information
- Scroll to zoom in/out of the network
- Download network as SVG for publications
Tool 3: Protein Relationship Analysis
This module analyzes the mutual exclusion and co-occurrence relationships between two specific proteins across samples, providing statistical significance and visual patterns of their association.
1 Input Configuration
Enter Two Proteins: Input Protein IDs (e.g., P12345) or Gene Names (e.g., TP53) for both proteins
Select Analysis Method:
- Fisher's Exact Test: Provides OR (Odds Ratio), P-value, FDR, and relationship type
- Jaccard Index: Provides Jaccard coefficient and relationship type
Choose Species: Human or Mouse
Select Tissue/Sample Type:
- Mouse: All, Liver, Brain
- Human: All, Tumor
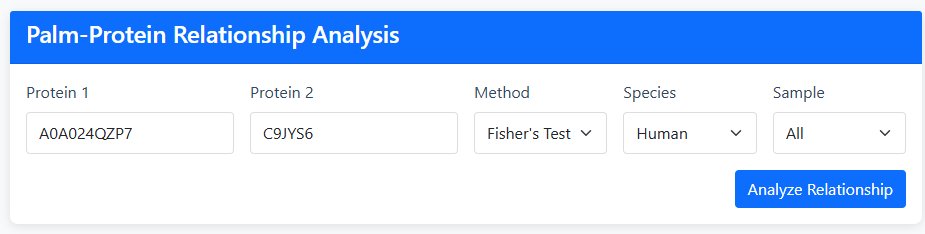
2 Analysis Execution
Click "Analyze Relationship" to execute the analysis and generate statistical results with visualization.
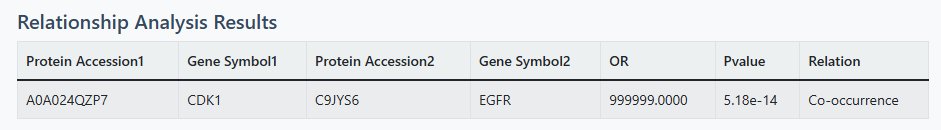
3 Results Interpretation
Statistical Results Table
- Protein Accession & Gene Symbol: Identifiers for both proteins
- Association Metrics:
- OR (Odds Ratio): Strength of association (>1 = positive, <1 = negative)
- P-value: Statistical significance of the relationship
- FDR: False Discovery Rate corrected P-value
- Jaccard: Similarity coefficient (0-1, higher = more similar)
- Relation Type:
- Co-occurrence (C): Proteins tend to appear together in samples
- Mutual Exclusion (M): Proteins rarely appear together in samples
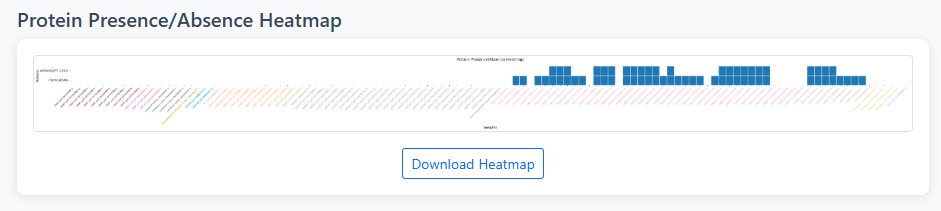
Heatmap Visualization
- Binary Representation: White (0) = protein absent, Blue (1) = protein present
- Sample Organization: Samples sorted alphabetically for easy comparison
- Tissue Color Coding: X-axis labels colored by tissue/cell type for quick identification
- Protein Labels: Display protein ID with gene symbol in parentheses when available
4 Interpretation Guide
Statistical Significance
- P-value < 0.05: Statistically significant relationship
- OR > 1: Positive association (co-occurrence tendency)
- OR < 1: Negative association (mutual exclusion tendency)
- Jaccard near 1: High expression pattern similarity
- Jaccard near 0: Low expression pattern similarity
Biological Interpretation
- Co-occurrence: May indicate functional cooperation, same pathway, or complex formation
- Mutual Exclusion: May indicate functional redundancy, different cellular states, or compensatory mechanisms
- Tissue-specific patterns: Reveal context-dependent relationships
Tool 4: Hotspot Mutation Analysis
Analyze the relationship between palmitoylation proteins and tumor hotspot mutations (CNVs/SNVs/genes) in human cell lines.
🔍 Search Modes
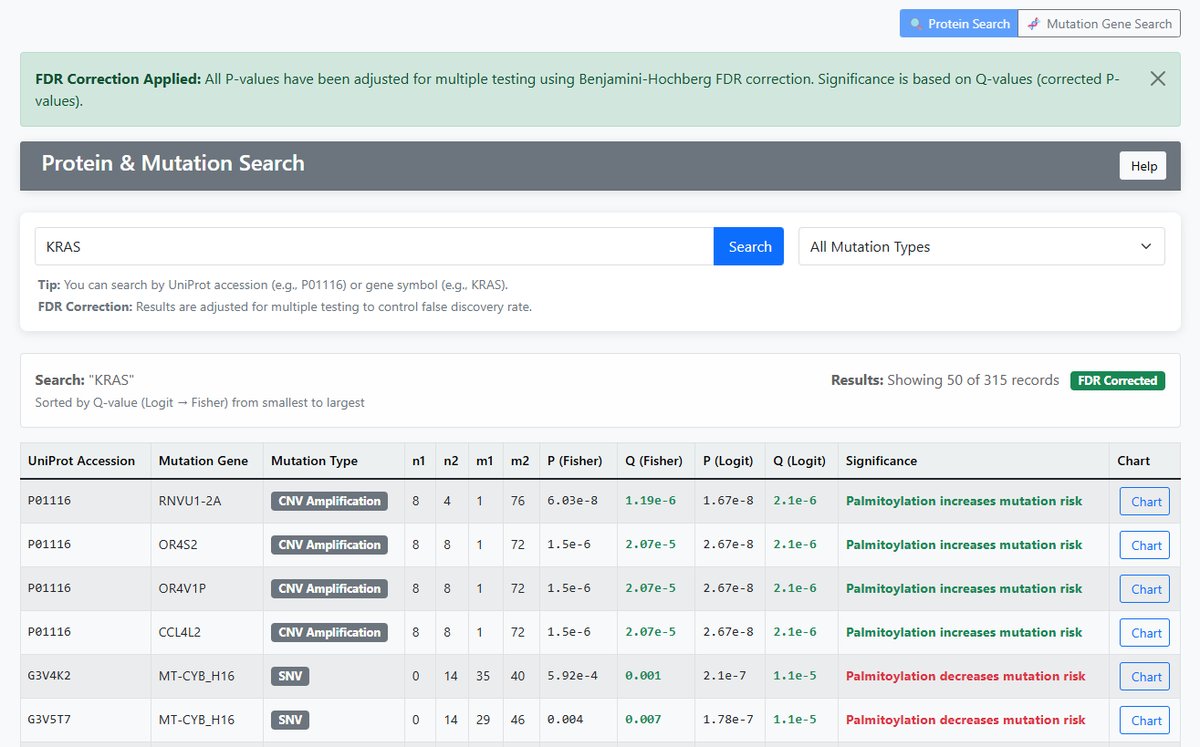
Protein Search
- Input: UniProt protein accession (e.g., P01116) or gene symbol (e.g., KRAS)
- Function: Search mutation information for specific proteins
- Output: All mutations associated with the protein
- Display Columns: Gene Symbol, Mutation Gene, Mutation Type
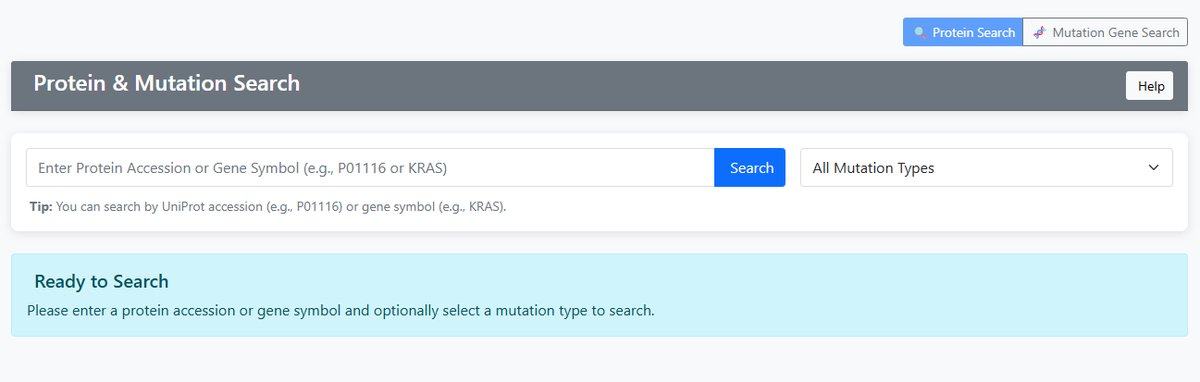
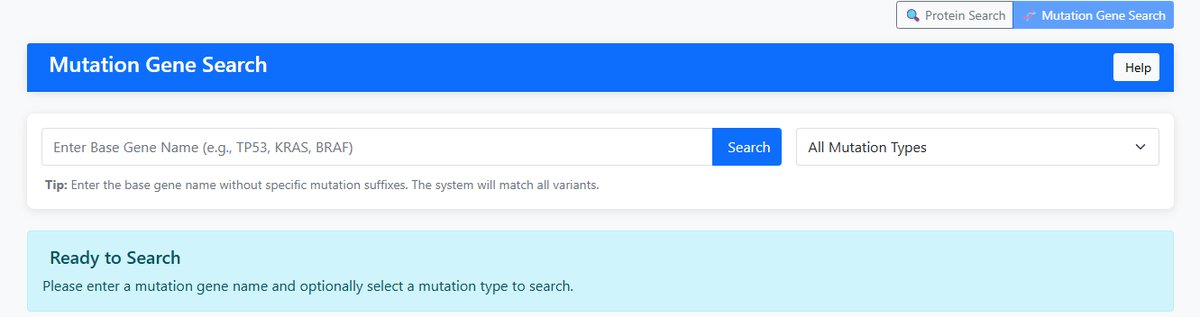
Mutation Gene Search
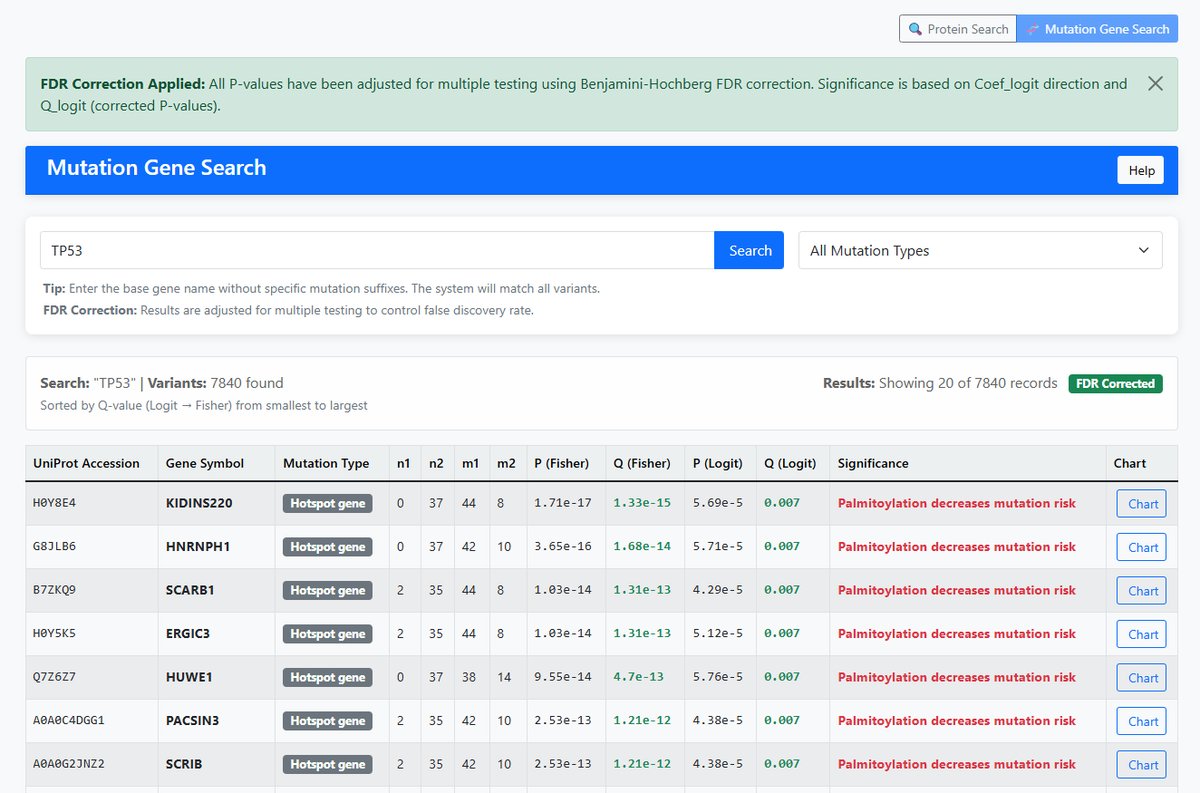
- Input: Base gene name (e.g., TP53, KRAS, BRAF)
- Function: Search all variants of a gene across different protein contexts
- Matching: Uses prefix matching (e.g., "TP53" matches "TP53_R273H", "TP53_M1863")
- Display Columns: UniProt Accession, Gene Symbol
📊 Mutation Types
CNV Amplification
- Definition: Copy number variation - gene amplification
- Impact: May lead to gene overexpression
- Significance: Associated with oncogene activation
CNV Deletion
- Definition: Copy number variation - gene deletion
- Impact: May lead to loss of gene function
- Significance: Associated with tumor suppressor gene inactivation
Hotspot Gene
- Definition: Genes with frequent mutations at specific positions
- Characteristics: Well-defined mutation hotspot regions
- Significance: Important markers of driver mutations
SNV
- Definition: Single nucleotide variations
- Types: Missense, nonsense, splice site mutations
- Impact: May affect protein structure and function
📈 Statistical Analysis
Statistical Methods
Fisher's Exact Test
- Purpose: Test association between mutation and palmitoylation
- Data: 2×2 contingency table counts
- Filtering: Results are filtered by adjust P (Fisher) < 0.05 and P (Logit) is available
Logit Regression
- Purpose: Model mutation probability based on palmitoylation
- Statistical Methods: Fisher's exact test and Firth logistic regression. The function for Firth logistic regression is “Mutation_status ~ Palmtoylation_status + Cell_line_backgound”
- Output: Coefficient (Coef_logit) and P-value
- Sorting: Results sorted by adjust P Logit and Fisher
FDR Correction
- Method: Benjamini-Hochberg FDR correction
- Output: adjust P (corrected P-values)
- Significance threshold: adjust P (Logit) < 0.05
- Display: Significant adjust P (Logit) shown in green
📋 Result Interpretation
Sample Count Columns
| Column | Description | Variable |
|---|---|---|
| n1 | Mutated and palmitoylated samples | mutated_palmitoylated |
| n2 | Mutated but non-palmitoylated samples | mutated_nonpalmitoylated |
| m1 | Wildtype but palmitoylated samples | wildtype_palmitoylated |
| m2 | Wildtype and non-palmitoylated samples | wildtype_nonpalmitoylated |
Statistical Columns
| Column | Description | Interpretation |
|---|---|---|
| P (Fisher) | Original Fisher's exact test P-value | Uncorrected significance level |
| adjust P (Fisher) | FDR-corrected Fisher's P-value | Multiple testing adjusted significance |
| P (Logit) | Original logistic regression P-value | Uncorrected significance level |
| adjust P (Logit) | FDR-corrected logistic regression P-value | Primary significance indicator |
Significance Interpretation
| Condition | Interpretation | Color Code |
|---|---|---|
| Coef_logit > 0 AND adjust P (Logit) < 0.05 | Palmitoylation increases mutation risk | Green |
| Coef_logit < 0 AND adjust P (Logit) < 0.05 | Palmitoylation decreases mutation risk | Red |
| adjust P (Logit) ≥ 0.05 | No significant association | Gray |
📊 Data Visualization
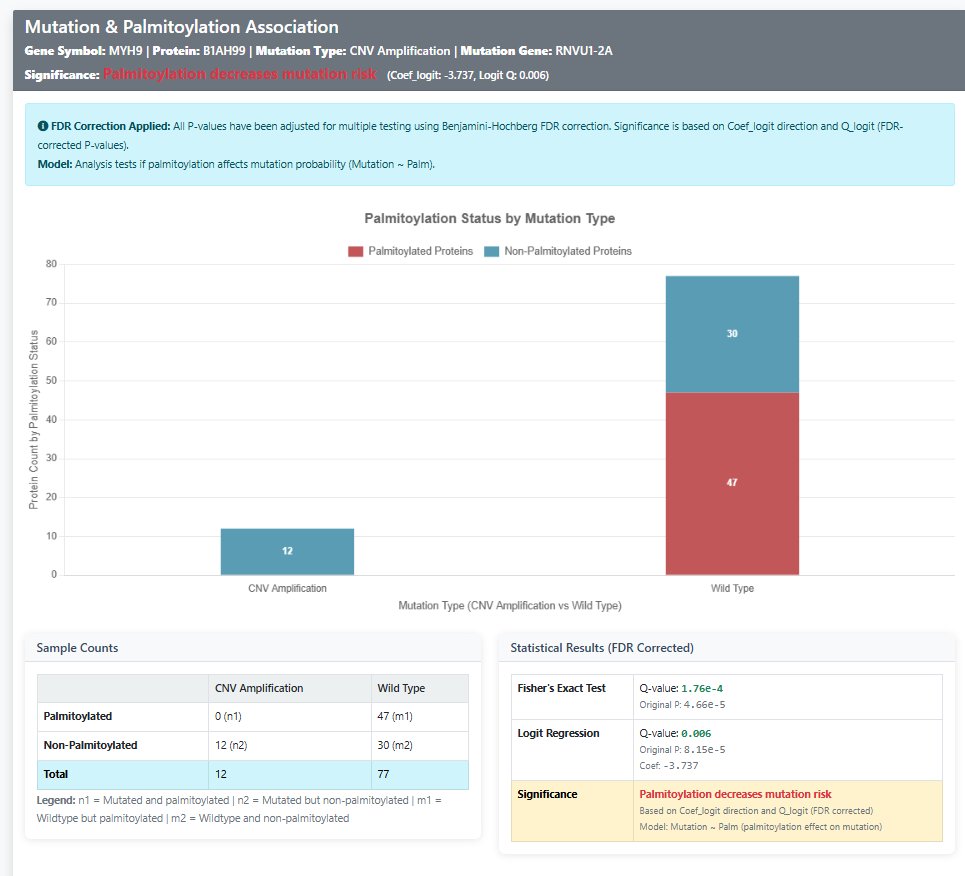
Interactive Bar Chart
- Access: Click "Chart" button in any result row
- Maroon Bars: Palmitoylated protein counts
- Blue Bars: Non-palmitoylated protein counts
- Comparison: Mutated vs wild-type samples side-by-side
- Data Labels: Exact count values displayed on bars
- Stacked Display: Clear visualization of palmitoylation status distribution
Statistical Panel
- Complete 2×2 contingency table with sample counts
- Both original P-values and FDR-corrected adjust P values
- Clear significance interpretation with color coding
- Regression coefficient with direction indication
- FDR correction status indicator
⚡ Performance Features
Optimization Techniques
- FDR Correction: Adjust P values are based on Benjamini-Hochberg FDR correction
- Caching: Frequently searched results cached for faster access (5-minute cache duration)
- Pagination: Large result sets efficiently paginated (15 records per page for protein search, 15 for mutation gene search)
- Smart Filtering: Results pre-filtered by by adjust P (Fisher) < 0.05 and P (Logit) is available
- Dynamic Column Display: Columns automatically shown/hidden based on search type
Search Tips
- Use base gene names for comprehensive variant searches (e.g., "TP53" instead of specific mutations)
- Filter by mutation type to focus on specific mutation categories
- Check adjust P (Logit) for reliable significance assessment (adjust P (Logit) indicates significance)
- Use the chart function for visual data exploration and better understanding of sample distributions
- Pay attention to color coding in results for quick significance assessment
Scientific Context
- Model: Analysis tests if palmitoylation affects mutation probability (Mutation ~ Palm)
- Biological Relevance: Understanding how post-translational modifications influence mutation patterns in cancer
- Clinical Significance: Identifying potential biomarkers and therapeutic targets
Tool 5: Multi-Protein Palmitoylation Pattern
Comprehensive analysis of the palmitoylation patterns of multiple proteins across different samples.
Go to Tool51 Input Configuration
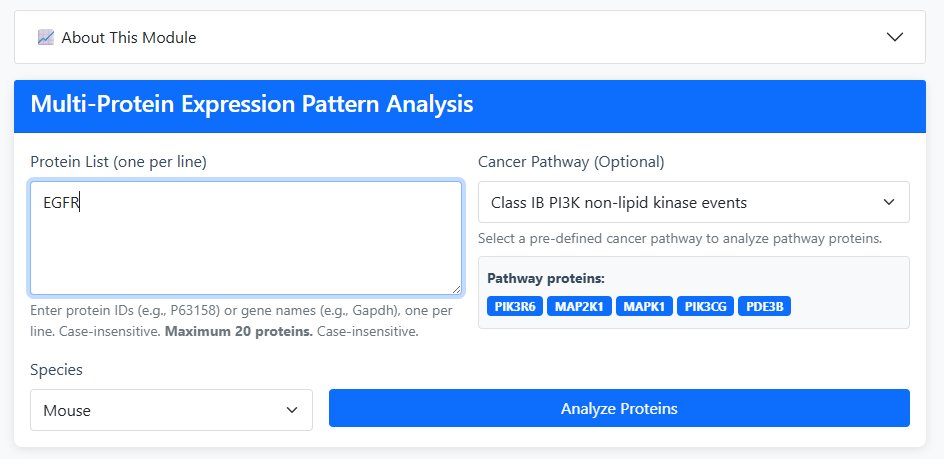
Input Method Selection:
- Manual Input: Enter protein IDs (e.g., P63158) or gene names (e.g., Gapdh) - one per line (maximum 20 proteins)
- Cancer Pathway Selection: Choose from pre-defined cancer signaling pathways (optional)
Select Species: Human or Mouse
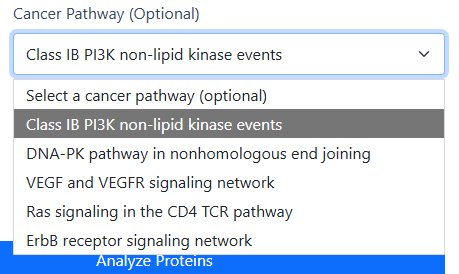
Available Cancer Pathways
- Class IB PI3K non-lipid kinase events
- DNA-PK pathway in nonhomologous end joining
- VEGF and VEGFR signaling network
- Ras signaling in the CD4 TCR pathway
- ErbB receptor signaling network
2 Analysis Execution
Click "Analyze Proteins" to execute the analysis and generate comprehensive visualizations.
3 Results Interpretation
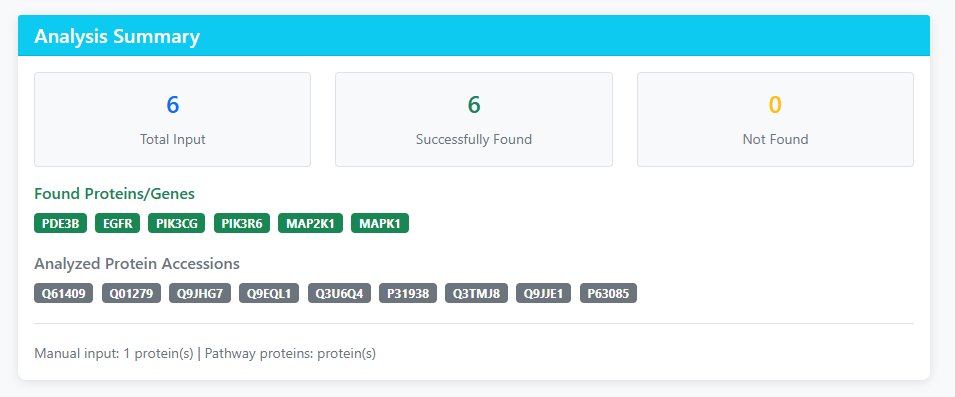
Analysis Summary
- Total Input: Number of proteins/gene names submitted
- Successfully Found: Proteins successfully matched in the database
- Not Found: Inputs that couldn't be matched (check spelling/species)
- Analyzed Protein Accessions: Actual protein IDs used in analysis
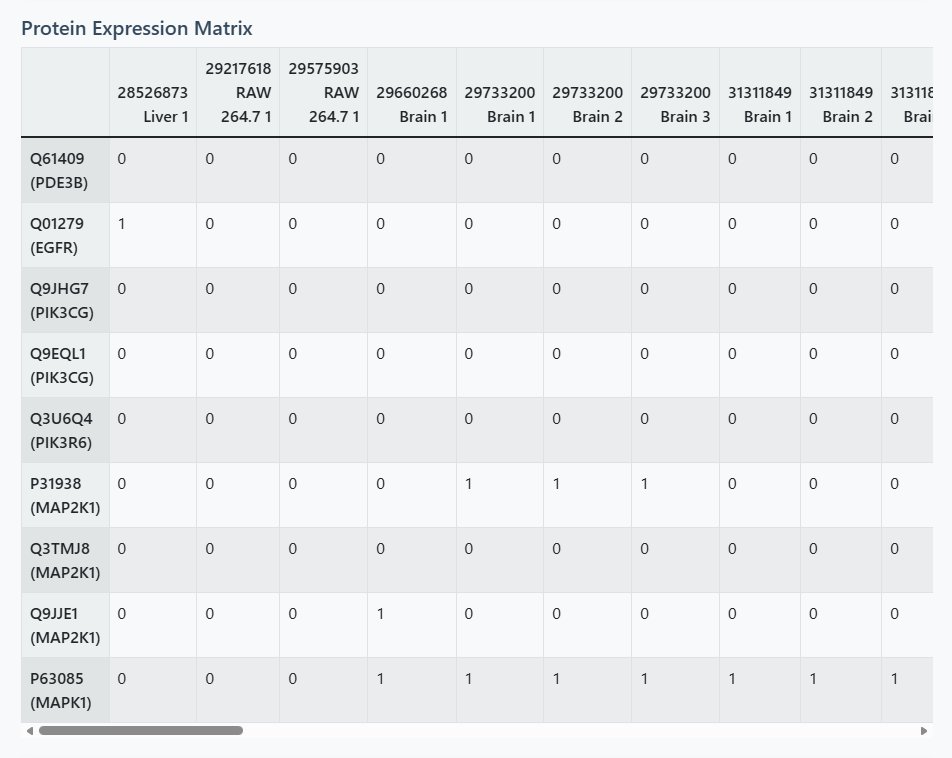
Heatmap Visualization
- Color Coding: White = absent (0), Blue = present (1)
- Dynamic Sizing: Automatically adjusts based on number of proteins and samples
- Tissue Color Labels: X-axis sample names colored by tissue/cell type
- Pattern Identification: Look for vertical patterns (sample clusters) and horizontal patterns (protein co-expression)
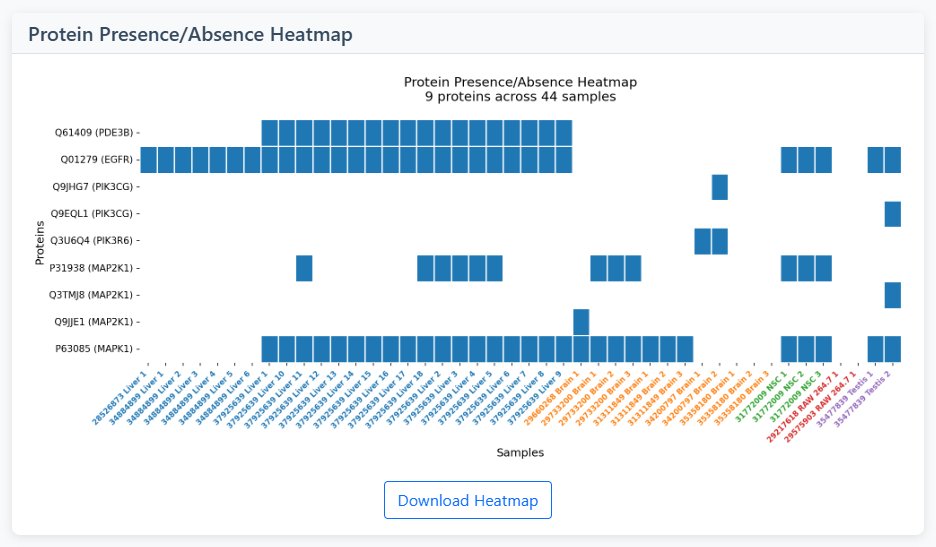
UMAP Visualization
- Dimensionality Reduction: Projects high-dimensional data into 2D space
- Sample Clustering: Similar samples cluster together in UMAP space
- Tissue Color Coding: Points colored by tissue/cell type
- Interpretation: Close clusters = similar expression patterns, distant points = distinct patterns
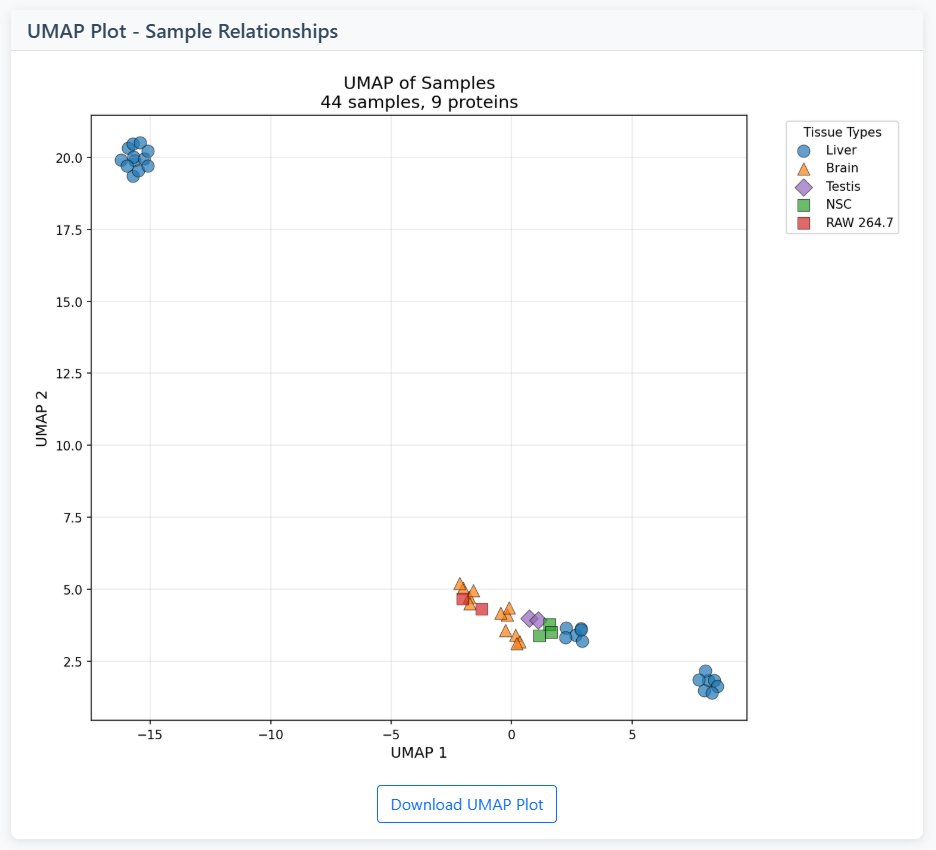
4 Interpretation Guide
Heatmap Patterns
- Vertical Stripes: Samples with similar protein expression profiles
- Horizontal Stripes: Proteins with similar expression across samples
- Block Patterns: Groups of proteins co-expressed in groups of samples
- Sparse Patterns: Proteins expressed in only a few samples
UMAP Patterns
- Tight Clusters: Samples with very similar expression patterns
- Separated Groups: Distinct sample types or conditions
- Gradient Patterns: Continuous variation in expression
- Outliers: Samples with unique expression profiles
Tool 6: Palmitoylation Motif Finder
Discover amino acid sequence motifs around palmitoylation sites in query amino acid sequences.
1 Input Configuration
Species Selection: Choose between Human or Mouse data
Protein Input: Enter UniProt accessions or gene symbols (maximum 100 proteins)
Data Sources: Select from three categories:
- Experimental: Experimentally verified sites from literature
- Database: Curated sites from SwissPalm, CysModDB, dbPTM, PTMD
- Prediction: Computationally predicted sites (High/Medium/Low confidence)
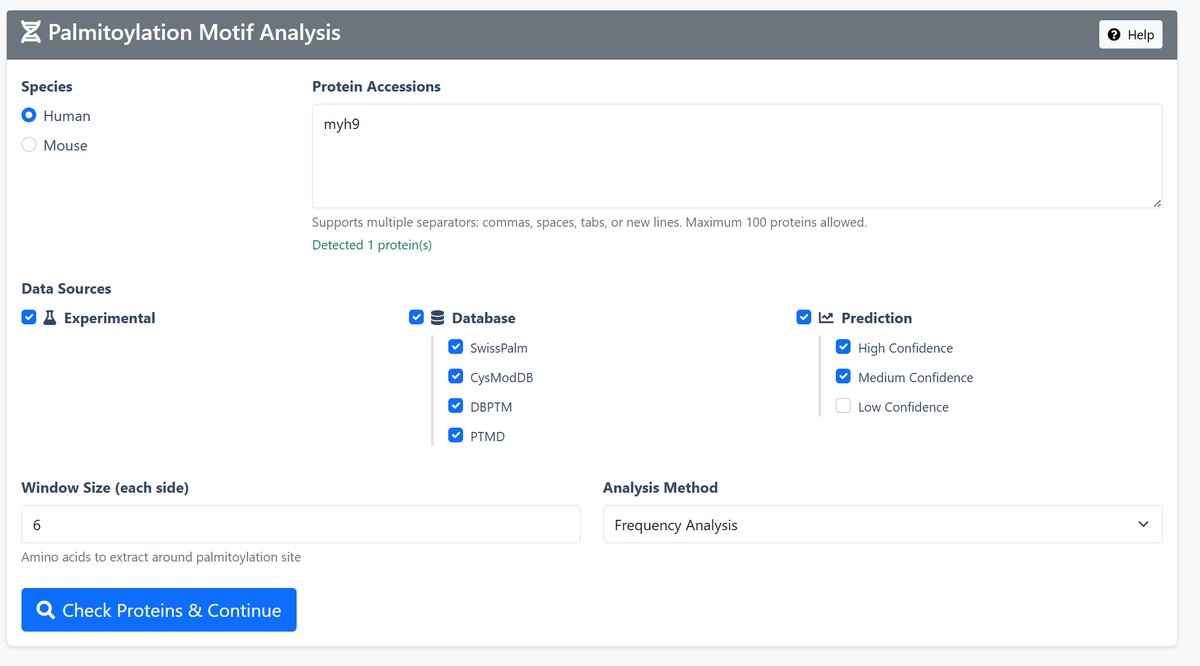
2 Analysis Parameters
Window Size: Number of amino acids to extract around each palmitoylation site (1-20, default: 6)
Analysis Method:
- Frequency Analysis: Basic amino acid frequency around sites
- Information Content: More sophisticated motif discovery
3 Protein Validation
After configuration, click "Check Proteins & Continue" to validate input proteins.
The validation page shows:
- Valid Proteins - Ready for analysis with available sites
- Proteins with Issues - Missing data or species mismatches
- Detailed breakdown of available data sources for each protein
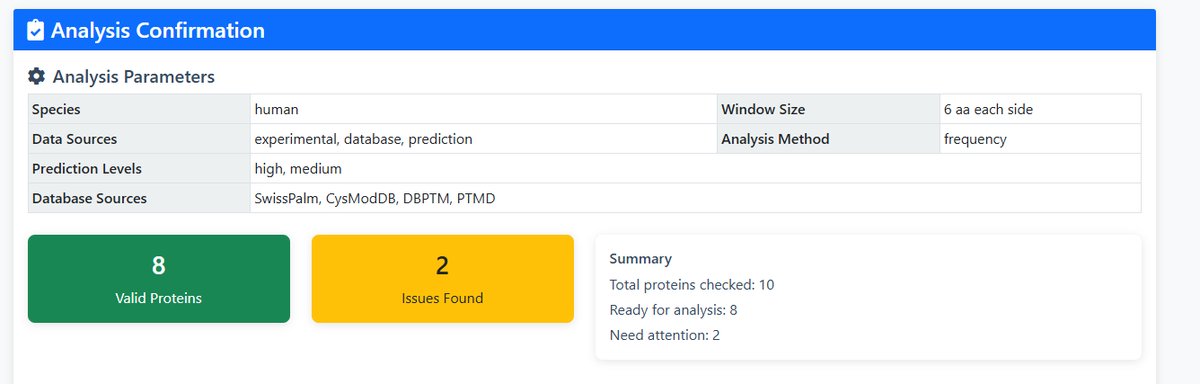
- This table provides basic information on palmitylated proteins suitable for further analysis
- Select the palmitylated proteins that you wish to analyse in order to proceed to the next step
- To view more detailed information about a specific palmitoylated protein, click 'Search' to navigate to the protein details page.
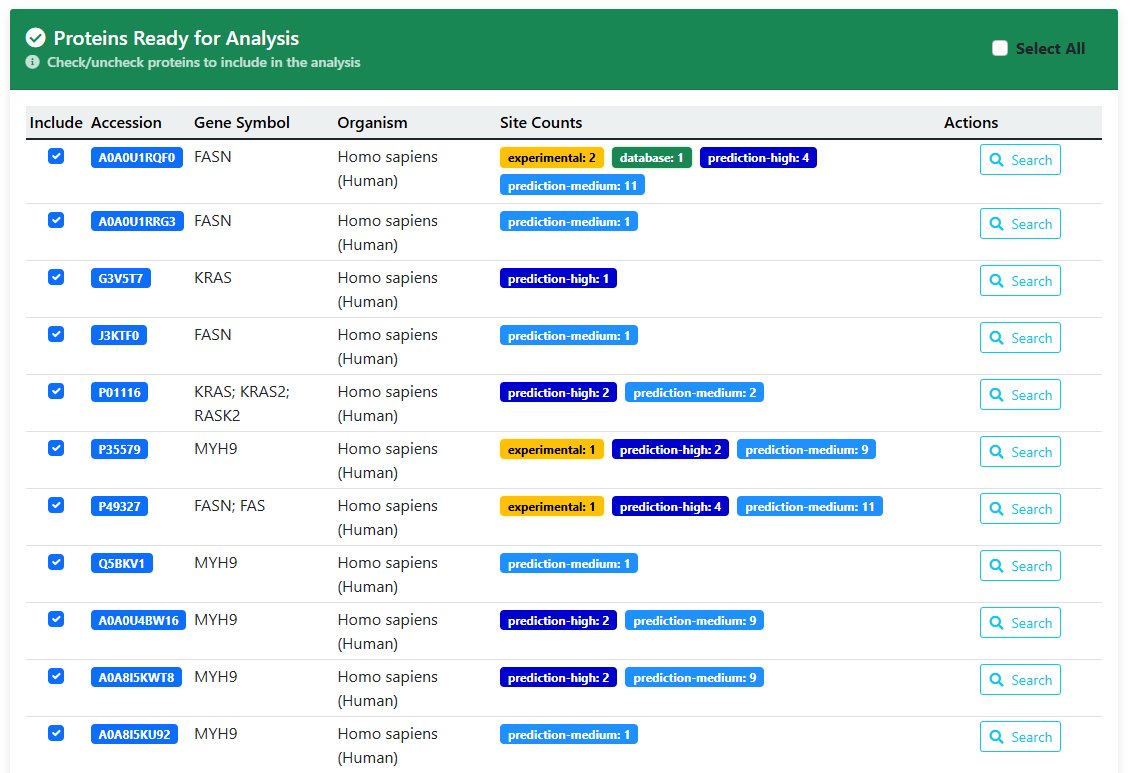
- This table provides basic information on palmitylated proteins that could not be analysed
- Reasons why proteins in the table cannot proceed to the next step of analysis
- Reason for error: The protein lacks a palmitoylation site; no related protein exists in the database.
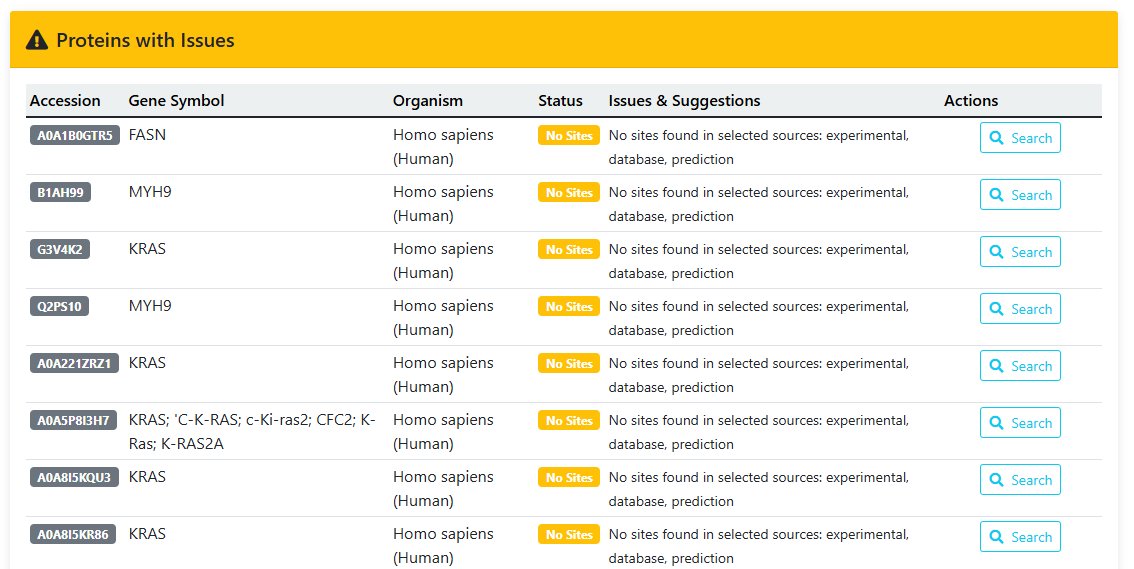
4 Analysis Execution
On the confirmation page, review parameters and click "Run Analysis" to start motif discovery.
5 Results Interpretation
The results page provides:
- Analysis Summary: Overview of proteins analyzed and motifs found
- Protein Details: List of analyzed proteins with site counts and positions
- Motif Logos: Sequence logos for discovered motifs, sorted by E-value
- Consensus Sequences: Representative sequences for each motif
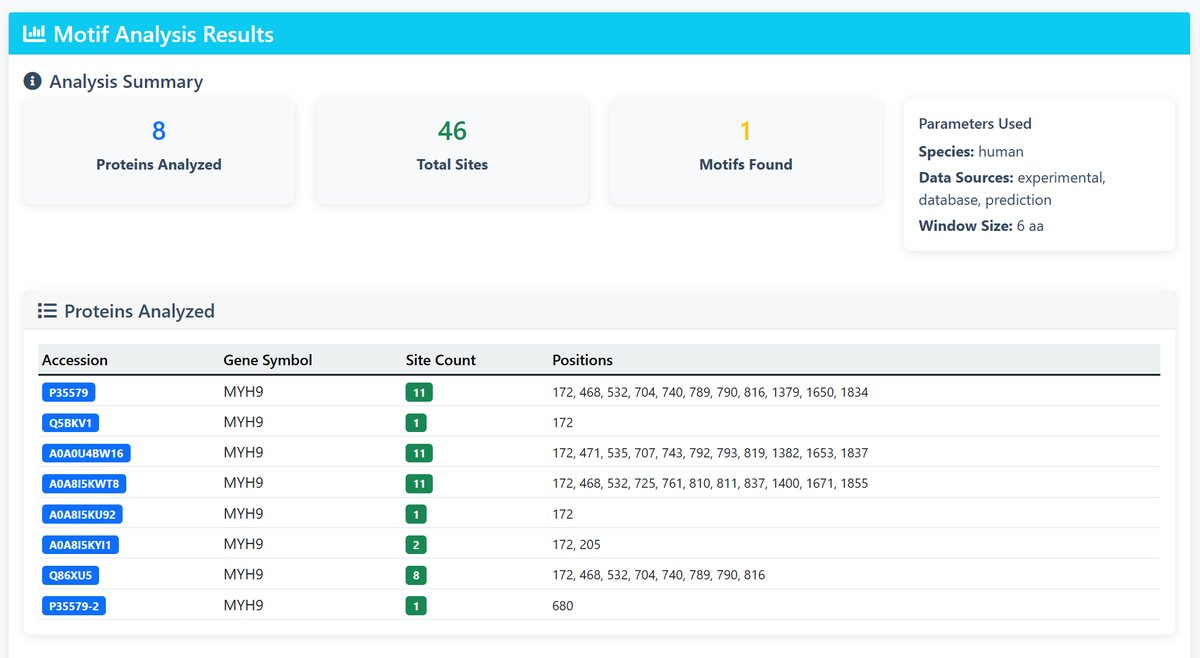
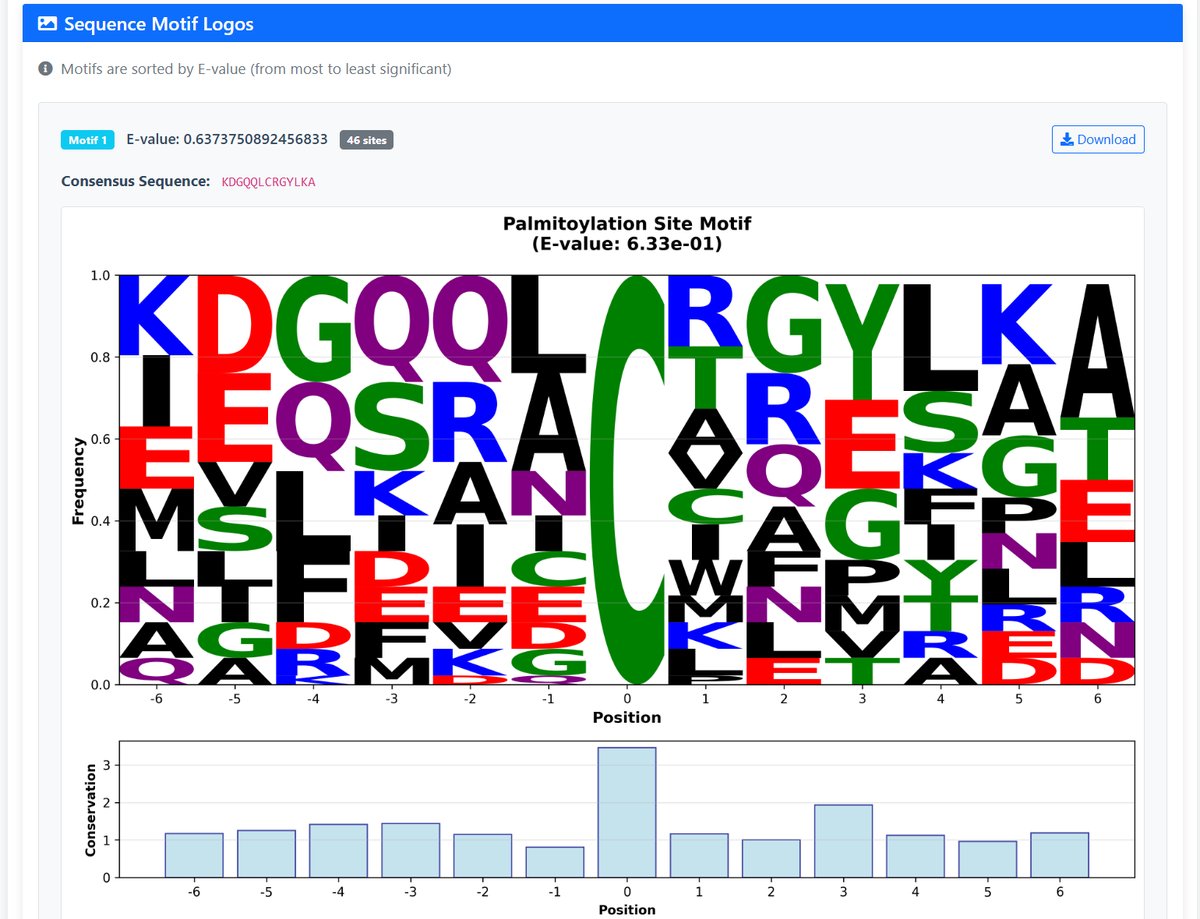
Understanding E-values: Lower E-values indicate more significant motifs (E-value < 0.05 is typically considered significant)
API Developer Interface
Programmatically access the Hotspot Mutation Analysis database with our RESTful API.
Quick Start
Base URL: https://palmlab.intelligent-oncology.com/api
All endpoints support GET requests with JSON responses by default
Jump to ExamplesComplete Python Tutorial
Download the comprehensive API usage tutorial with working examples:
Download api_usage_tutorial.pyIncludes search functions, pagination handling, and data analysis examples for all tools.
API Overview
- Base URL:
https://palmlab.intelligent-oncology.com/api - HTTP Method: GET for all endpoints
- Response Formats: JSON (default), CSV, PNG (for visualization endpoints)
- Features: Pagination, Format selection, Image generation, Batch processing
Available Endpoints
| Tool | Endpoint | Description | Required Parameters | Default Values |
|---|---|---|---|---|
| Tools1 | /tools1/differential/ |
Differential expression analysis | proteins (comma-separated list) |
species=human, format=json |
| Tools2 | /tools2/network/ |
Protein interaction partners (shows top 50) | protein (single protein) |
species=Mouse, tissue=All, format=json |
| Tools3 | /tools3/pair/ |
Protein pair relationship | protein1, protein2 |
species=Mouse, tissue=All, format=json |
| Tools4a | /tools4/searchpalmitoylatedprotein/ |
Search protein mutations (human only) | protein_query (accession or gene symbol) |
mutation_type_choice=All, page=1, page_size=20 |
| Tools4b | /tools4/searchgenemutation/ |
Search gene mutations (base gene name matching) | mutation_gene_query |
mutation_type_choice=All, page=1, page_size=20 |
| Tools5 | /tools5/multi/ |
Multi-protein expression analysis | proteins (comma-separated list) |
species=Mouse, format=json, include_pca=false |
| Tools6 | /tools6/motif/ |
Motif pattern analysis | proteins (comma-separated list) |
species=human, format=image, window_size=6, data_sources=experimental,database,prediction |
Parameter Details
Common Parameters:
-
format- Response format
Values:json(default),csv,image(for tools with visualization) -
download- Force file download
Values:trueorfalse(default:false) -
species- Organism
Values:Human,Mouse
Tool-Specific Parameters:
-
Tools4:
mutation_type_choice
Values:CNV_Amplification,CNV_Deletion,Hotspot_gene,SNV,All(default) -
Tools5:
include_pca
Values:trueorfalse(default:false) -
Tools6:
data_sources
Values:experimental,database,prediction(comma-separated, default: all)
API Examples
Tools1: Differential Expression Analysis
Multiple modes supported:
- Default mode: For human: cancer vs normal; For mouse: auto-grouping
- Cancer vs Normal mode: All cancer datasets vs all normal datasets (human only)
- Custom mode: User-defined group A vs group B (both human and mouse)
# ==================== MODE 1: Default (species-based) ====================
# For human: auto cancer vs normal; For mouse: auto grouping
curl "https://palmlab.intelligent-oncology.com/api/tools1/differential/?proteins=P01112,P07900,A0A0J9YXG8,P17252,P78460,A0A169TED2,L7RSM7,P01135,P05771&species=human"
# ==================== MODE 2: Cancer vs Normal (human only) ====================
curl "https://palmlab.intelligent-oncology.com/api/tools1/differential/?proteins=P01112,P07900,A0A0J9YXG8,P17252,P78460,A0A169TED2,L7RSM7,P01135,P05771&species=human&mode=cancer_vs_normal"
# ==================== MODE 3: Custom Group A vs Group B ====================
# Human example: custom cancer vs normal comparison
curl "https://palmlab.intelligent-oncology.com/api/tools1/differential/?proteins=P01112,P07900,A0A0J9YXG8,P17252,P78460,A0A169TED2,L7RSM7,P01135,P05771&species=human&mode=custom&group_a_datasets=Jurkat_T_cells,LNCaP_cells&group_b_datasets=T_cells,293T_cells&group_a_label=Cancer&group_b_label=Normal"
# Mouse example: custom tissue comparison
curl "https://palmlab.intelligent-oncology.com/api/tools1/differential/?proteins=Q3U6Q4,Q61409&species=mouse&mode=custom&group_a_datasets=brain_tissue,liver_tissue&group_b_datasets=testis,Macrophage_Raw_264.7&group_a_label=Brain_Liver&group_b_label=Testis_Macrophage"
# ==================== Format and Download Options ====================
# Download as CSV
curl "https://palmlab.intelligent-oncology.com/api/tools1/differential/?proteins=P01112,P07900,A0A0J9YXG8,P17252,P78460,A0A169TED2,L7RSM7,P01135,P05771&mode=cancer_vs_normal&format=csv&download=true" -o differential.csv
# Get results as JSON (default)
curl "https://palmlab.intelligent-oncology.com/api/tools1/differential/?proteins=P01112,P07900,A0A0J9YXG8,P17252,P78460,A0A169TED2,L7RSM7,P01135,P05771&species=human&mode=default"
# ==================== Get Available Datasets ====================
# Get all human datasets for cancer vs normal mode
curl "https://palmlab.intelligent-oncology.com/api/tools1/datasets/?species=human&analysis_type=cancer_vs_normal"
# Get all mouse datasets
curl "https://palmlab.intelligent-oncology.com/api/tools1/datasets/?species=mouse"
# Get all human datasets (mixed)
curl "https://palmlab.intelligent-oncology.com/api/tools1/datasets/?species=human&analysis_type=all"Tools2: Protein Interaction Network
Note: Shows top 50 interactions sorted by Studies Co-occur and Odds Ratio.
# Find interaction partners for P19096 (Mouse, all tissues)
curl "https://palmlab.intelligent-oncology.com/api/tools2/network/?protein=P19096&species=Mouse&tissue=All"
# Human protein with tumor tissue
curl "https://palmlab.intelligent-oncology.com/api/tools2/network/?protein=P01112&species=Human&tissue=Tumor"
# Get results in CSV format
curl "https://palmlab.intelligent-oncology.com/api/tools2/network/?protein=P19096&species=Mouse&format=csv&download=true" -o interactions.csvTools3: Protein Pair Analysis
# Analyze relationship between P19096 and Q01279
curl "https://palmlab.intelligent-oncology.com/api/tools3/pair/?protein1=P19096&protein2=Q01279&species=Mouse"
# With heatmap image (PNG format)
curl "https://palmlab.intelligent-oncology.com/api/tools3/pair/?protein1=P19096&protein2=Q01279&output_image=true&format=image&download=true" -o pair_heatmap.png
Tools4: Mutation Analysis
Important: Uses FDR correction and matches web interface format.
Tools4a: Search by Protein
Searches human proteins only by accession or exact gene symbol match.
# Search mutations in KRAS protein (P01116) - all mutation types
curl "https://palmlab.intelligent-oncology.com/api/tools4/searchpalmitoylatedprotein/?protein_query=P01116"
# Search by gene symbol (exact match)
curl "https://palmlab.intelligent-oncology.com/api/tools4/searchpalmitoylatedprotein/?protein_query=KRAS"
# Filter by SNV mutations only
curl "https://palmlab.intelligent-oncology.com/api/tools4/searchpalmitoylatedprotein/?protein_query=P01116&mutation_type_choice=SNV&page=2&page_size=15"
# Download as CSV (matches web interface columns)
curl "https://palmlab.intelligent-oncology.com/api/tools4/searchpalmitoylatedprotein/?protein_query=P01116&format=csv&download=true" -o kras_mutations.csvTools4b: Search by Gene Mutation
Uses base gene name matching (e.g., "TP53" matches "TP53", "TP53-AS1", etc.).
# Search TP53 gene mutations
curl "https://palmlab.intelligent-oncology.com/api/tools4/searchgenemutation/?mutation_gene_query=TP53"
# Filter by Hotspot Gene
curl "https://palmlab.intelligent-oncology.com/api/tools4/searchgenemutation/?mutation_gene_query=TP53&mutation_type_choice=Hotspot_gene"
# Get CSV format
curl "https://palmlab.intelligent-oncology.com/api/tools4/searchgenemutation/?mutation_gene_query=TP53&format=csv&download=true" -o tp53_mutations.csvTools5: Multi-Protein Analysis
# Analyze multiple proteins (Mouse)
curl "https://palmlab.intelligent-oncology.com/api/tools5/multi/?proteins=Q3U6Q4,Q61409,Q9EQL1,P63085&species=Mouse"
# Get heatmap as PNG
curl "https://palmlab.intelligent-oncology.com/api/tools5/multi/?proteins=Q3U6Q4,Q61409,Q9EQL1,P63085&output_image=true&format=image&download=true" -o expression_heatmap.png
# Get PCA image only
curl "https://palmlab.intelligent-oncology.com/api/tools5/multi/?proteins=Q3U6Q4,Q61409,Q9EQL1,P63085&output_image=true&include_pca=true&format=image&download=true" -o expression_pca.pngTools6: Motif Analysis
Note: Returns PNG image by default.
# Analyze motifs in multiple proteins (returns PNG)
curl "https://palmlab.intelligent-oncology.com/api/tools6/motif/?proteins=P01112,P07900,A0A0J9YXG8,P17252,P78460&species=human" -o motif.png
# Get JSON response instead
curl "https://palmlab.intelligent-oncology.com/api/tools6/motif/?proteins=P01112,P07900&species=human&format=json"
# Customize analysis parameters
curl "https://palmlab.intelligent-oncology.com/api/tools6/motif/?proteins=P01112,P07900&species=human&window_size=8&data_sources=experimental,database&analysis_method=information"
# Download as CSV summary
curl "https://palmlab.intelligent-oncology.com/api/tools6/motif/?proteins=P01112,P07900&format=csv&download=true" -o motif_analysis.csvResponse Format
All endpoints return consistent JSON structure when format=json:
{
"success": true,
"message": "Analysis completed successfully",
"query": {
"proteins": ["P01112", "P07900"],
"species": "human",
"format": "json",
"timestamp": "2024-01-15T10:30:00Z"
},
"results": [
{
"protein_accession": "P01112",
"gene_symbol": "HRAS",
"cancer_ratio": 0.75,
"normal_ratio": 0.25,
"odds_ratio": 9.0,
"p_value": 0.001,
"significant": true
}
],
"summary": {
"total_found": 1,
"significant_count": 1,
"note": "Showing top 50 results"
}
}Common Response Fields:
success- Boolean indicating request successmessage- Human-readable message in Englishquery- Parameters used for the requestresults- Array of analysis resultssummary- Statistical summary and notes
Error Response:
{
"success": false,
"message": "Protein not found: XYZ123",
"error": "Invalid protein ID",
"status": 400,
"timestamp": "2024-01-15T10:30:00Z"
}